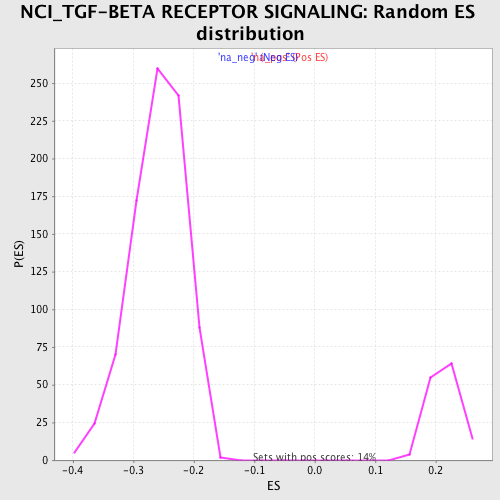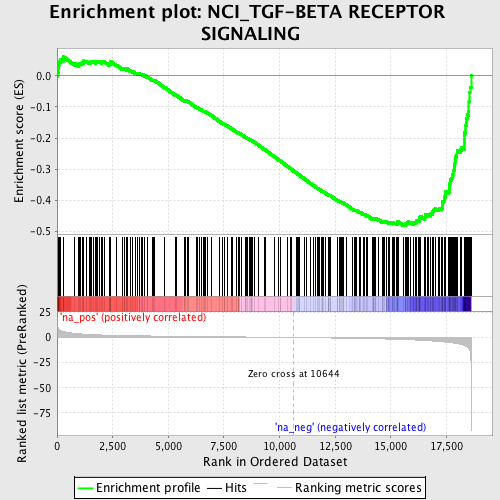
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_truncNotch_versus_normalThy |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | NCI_TGF-BETA RECEPTOR SIGNALING |
| Enrichment Score (ES) | -0.48339587 |
| Normalized Enrichment Score (NES) | -1.8674273 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.06799437 |
| FWER p-Value | 0.672 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CEBPB | 39 | 9.502 | 0.0113 | No | ||
| 2 | IL5 | 45 | 9.359 | 0.0243 | No | ||
| 3 | MAPKAPK2 | 58 | 8.591 | 0.0358 | No | ||
| 4 | CDK4 | 86 | 7.778 | 0.0454 | No | ||
| 5 | FGR | 143 | 7.022 | 0.0523 | No | ||
| 6 | EIF4EBP1 | 268 | 6.109 | 0.0542 | No | ||
| 7 | EEF2K | 289 | 5.954 | 0.0615 | No | ||
| 8 | BAD | 784 | 4.120 | 0.0405 | No | ||
| 9 | PIAS3 | 979 | 3.746 | 0.0352 | No | ||
| 10 | HSPB1 | 990 | 3.722 | 0.0399 | No | ||
| 11 | EGR3 | 1068 | 3.569 | 0.0408 | No | ||
| 12 | IL2RA | 1123 | 3.488 | 0.0428 | No | ||
| 13 | MYC | 1180 | 3.415 | 0.0446 | No | ||
| 14 | MAP3K7IP1 | 1199 | 3.378 | 0.0484 | No | ||
| 15 | BLK | 1301 | 3.219 | 0.0475 | No | ||
| 16 | USF1 | 1464 | 3.021 | 0.0429 | No | ||
| 17 | CDC25B | 1519 | 2.963 | 0.0442 | No | ||
| 18 | ARRB2 | 1536 | 2.944 | 0.0475 | No | ||
| 19 | RBBP4 | 1624 | 2.853 | 0.0468 | No | ||
| 20 | TP53 | 1734 | 2.750 | 0.0447 | No | ||
| 21 | DAB2 | 1770 | 2.714 | 0.0467 | No | ||
| 22 | CDKN1A | 1821 | 2.661 | 0.0477 | No | ||
| 23 | PPP1CA | 1885 | 2.600 | 0.0480 | No | ||
| 24 | TCF3 | 2016 | 2.484 | 0.0444 | No | ||
| 25 | DUSP10 | 2057 | 2.463 | 0.0457 | No | ||
| 26 | PRKACA | 2114 | 2.419 | 0.0461 | No | ||
| 27 | IL4 | 2334 | 2.280 | 0.0374 | No | ||
| 28 | MAP3K10 | 2364 | 2.261 | 0.0390 | No | ||
| 29 | FKBP8 | 2409 | 2.232 | 0.0398 | No | ||
| 30 | SHC1 | 2417 | 2.229 | 0.0426 | No | ||
| 31 | RUNX3 | 2418 | 2.228 | 0.0457 | No | ||
| 32 | SLC3A2 | 2671 | 2.093 | 0.0350 | No | ||
| 33 | MAPK3 | 2925 | 1.958 | 0.0240 | No | ||
| 34 | RBBP7 | 3007 | 1.925 | 0.0223 | No | ||
| 35 | ITGB5 | 3044 | 1.908 | 0.0230 | No | ||
| 36 | IL10 | 3140 | 1.863 | 0.0205 | No | ||
| 37 | RNF128 | 3148 | 1.861 | 0.0227 | No | ||
| 38 | YWHAQ | 3313 | 1.791 | 0.0163 | No | ||
| 39 | PKN1 | 3396 | 1.761 | 0.0144 | No | ||
| 40 | GRB2 | 3538 | 1.714 | 0.0091 | No | ||
| 41 | CTDSP1 | 3599 | 1.690 | 0.0082 | No | ||
| 42 | RPS6KA4 | 3689 | 1.664 | 0.0058 | No | ||
| 43 | RUNX2 | 3706 | 1.658 | 0.0072 | No | ||
| 44 | COL1A2 | 3772 | 1.638 | 0.0060 | No | ||
| 45 | SERPINE1 | 3851 | 1.612 | 0.0040 | No | ||
| 46 | CALM1 | 3948 | 1.579 | 0.0011 | No | ||
| 47 | E2F1 | 4079 | 1.545 | -0.0038 | No | ||
| 48 | ETV1 | 4288 | 1.484 | -0.0131 | No | ||
| 49 | TGIF2 | 4345 | 1.467 | -0.0140 | No | ||
| 50 | NKX2-5 | 4390 | 1.455 | -0.0144 | No | ||
| 51 | EIF4E | 4842 | 1.338 | -0.0370 | No | ||
| 52 | ESR1 | 5304 | 1.220 | -0.0604 | No | ||
| 53 | LYN | 5368 | 1.203 | -0.0622 | No | ||
| 54 | CDKN2B | 5729 | 1.115 | -0.0802 | No | ||
| 55 | TBX21 | 5746 | 1.112 | -0.0795 | No | ||
| 56 | ATF1 | 5760 | 1.109 | -0.0786 | No | ||
| 57 | IFNB1 | 5849 | 1.085 | -0.0819 | No | ||
| 58 | PRKCZ | 5893 | 1.076 | -0.0827 | No | ||
| 59 | ATF3 | 6265 | 0.994 | -0.1015 | No | ||
| 60 | SNTA1 | 6327 | 0.979 | -0.1034 | No | ||
| 61 | MAP3K6 | 6399 | 0.965 | -0.1059 | No | ||
| 62 | CAV1 | 6478 | 0.945 | -0.1088 | No | ||
| 63 | MAPK13 | 6583 | 0.919 | -0.1132 | No | ||
| 64 | CTGF | 6609 | 0.915 | -0.1133 | No | ||
| 65 | DLX1 | 6672 | 0.896 | -0.1154 | No | ||
| 66 | ZBTB17 | 6780 | 0.872 | -0.1200 | No | ||
| 67 | TAOK2 | 6927 | 0.840 | -0.1267 | No | ||
| 68 | PARD6A | 7311 | 0.759 | -0.1465 | No | ||
| 69 | SAP18 | 7432 | 0.735 | -0.1520 | No | ||
| 70 | PTPN1 | 7537 | 0.714 | -0.1567 | No | ||
| 71 | MAPKAPK5 | 7540 | 0.713 | -0.1558 | No | ||
| 72 | PRKCA | 7649 | 0.691 | -0.1607 | No | ||
| 73 | XPO1 | 7829 | 0.653 | -0.1695 | No | ||
| 74 | HCK | 7871 | 0.643 | -0.1708 | No | ||
| 75 | RAN | 8046 | 0.606 | -0.1794 | No | ||
| 76 | TH | 8138 | 0.583 | -0.1836 | No | ||
| 77 | CCND1 | 8140 | 0.583 | -0.1828 | No | ||
| 78 | NR4A1 | 8191 | 0.573 | -0.1847 | No | ||
| 79 | VDR | 8273 | 0.557 | -0.1883 | No | ||
| 80 | SOS1 | 8457 | 0.519 | -0.1976 | No | ||
| 81 | CITED1 | 8487 | 0.511 | -0.1984 | No | ||
| 82 | DUSP8 | 8517 | 0.504 | -0.1993 | No | ||
| 83 | MAPK12 | 8571 | 0.491 | -0.2015 | No | ||
| 84 | MAP3K4 | 8643 | 0.476 | -0.2047 | No | ||
| 85 | AKT1 | 8696 | 0.467 | -0.2068 | No | ||
| 86 | PPARG | 8728 | 0.461 | -0.2079 | No | ||
| 87 | NR3C1 | 8735 | 0.461 | -0.2075 | No | ||
| 88 | HNF4A | 8772 | 0.454 | -0.2089 | No | ||
| 89 | CTBP1 | 8868 | 0.429 | -0.2134 | No | ||
| 90 | IL2 | 9059 | 0.390 | -0.2232 | No | ||
| 91 | CSF2 | 9327 | 0.329 | -0.2373 | No | ||
| 92 | GADD45G | 9370 | 0.321 | -0.2391 | No | ||
| 93 | SMAD7 | 9750 | 0.233 | -0.2594 | No | ||
| 94 | CASP3 | 9967 | 0.184 | -0.2709 | No | ||
| 95 | PRKCE | 10026 | 0.168 | -0.2739 | No | ||
| 96 | PAK3 | 10035 | 0.165 | -0.2741 | No | ||
| 97 | RIPK1 | 10351 | 0.086 | -0.2911 | No | ||
| 98 | MAP2K3 | 10472 | 0.051 | -0.2976 | No | ||
| 99 | PPARGC1A | 10520 | 0.038 | -0.3001 | No | ||
| 100 | FOS | 10770 | -0.029 | -0.3136 | No | ||
| 101 | IL3 | 10817 | -0.043 | -0.3160 | No | ||
| 102 | TNF | 10851 | -0.051 | -0.3178 | No | ||
| 103 | YWHAZ | 10875 | -0.059 | -0.3189 | No | ||
| 104 | EGR4 | 10880 | -0.060 | -0.3191 | No | ||
| 105 | PRKCH | 11098 | -0.114 | -0.3307 | No | ||
| 106 | AR | 11203 | -0.146 | -0.3362 | No | ||
| 107 | STMN1 | 11411 | -0.206 | -0.3472 | No | ||
| 108 | SKIL | 11504 | -0.235 | -0.3518 | No | ||
| 109 | FOSL1 | 11626 | -0.268 | -0.3581 | No | ||
| 110 | YES1 | 11724 | -0.298 | -0.3629 | No | ||
| 111 | MAPKAPK3 | 11729 | -0.299 | -0.3627 | No | ||
| 112 | CAMK2B | 11809 | -0.327 | -0.3666 | No | ||
| 113 | FOXH1 | 11896 | -0.349 | -0.3707 | No | ||
| 114 | PTGS2 | 11918 | -0.354 | -0.3714 | No | ||
| 115 | STRAP | 11928 | -0.360 | -0.3714 | No | ||
| 116 | MYOD1 | 11963 | -0.369 | -0.3727 | No | ||
| 117 | GSC | 12061 | -0.397 | -0.3774 | No | ||
| 118 | CD40LG | 12191 | -0.434 | -0.3838 | No | ||
| 119 | PRKCD | 12195 | -0.435 | -0.3834 | No | ||
| 120 | MEF2C | 12220 | -0.444 | -0.3840 | No | ||
| 121 | NFATC2 | 12258 | -0.453 | -0.3854 | No | ||
| 122 | BAMBI | 12283 | -0.460 | -0.3861 | No | ||
| 123 | ZFYVE9 | 12294 | -0.465 | -0.3860 | No | ||
| 124 | YWHAE | 12605 | -0.561 | -0.4021 | No | ||
| 125 | RPS6KA5 | 12623 | -0.567 | -0.4022 | No | ||
| 126 | TFDP1 | 12684 | -0.584 | -0.4046 | No | ||
| 127 | FOXO3 | 12713 | -0.594 | -0.4053 | No | ||
| 128 | JUN | 12736 | -0.603 | -0.4056 | No | ||
| 129 | OCLN | 12781 | -0.621 | -0.4072 | No | ||
| 130 | SKI | 12814 | -0.633 | -0.4080 | No | ||
| 131 | DACT2 | 12854 | -0.644 | -0.4092 | No | ||
| 132 | NUP153 | 12857 | -0.645 | -0.4084 | No | ||
| 133 | PLA2G4A | 13012 | -0.699 | -0.4158 | No | ||
| 134 | CAMK2A | 13297 | -0.813 | -0.4301 | No | ||
| 135 | FOXG1 | 13372 | -0.842 | -0.4330 | No | ||
| 136 | MEF2D | 13381 | -0.847 | -0.4322 | No | ||
| 137 | RAC1 | 13414 | -0.858 | -0.4327 | No | ||
| 138 | IFNG | 13438 | -0.868 | -0.4328 | No | ||
| 139 | SRC | 13590 | -0.923 | -0.4397 | No | ||
| 140 | MAP3K5 | 13604 | -0.929 | -0.4391 | No | ||
| 141 | RAB5A | 13643 | -0.942 | -0.4398 | No | ||
| 142 | ELK4 | 13777 | -1.001 | -0.4456 | No | ||
| 143 | TSC2 | 13804 | -1.010 | -0.4456 | No | ||
| 144 | MAP3K7 | 13920 | -1.056 | -0.4504 | No | ||
| 145 | DYNLRB1 | 13962 | -1.073 | -0.4511 | No | ||
| 146 | ZFYVE16 | 14180 | -1.177 | -0.4612 | No | ||
| 147 | YAP1 | 14203 | -1.189 | -0.4608 | No | ||
| 148 | GADD45B | 14233 | -1.204 | -0.4606 | No | ||
| 149 | SNIP1 | 14242 | -1.207 | -0.4594 | No | ||
| 150 | PRKG1 | 14283 | -1.226 | -0.4598 | No | ||
| 151 | MAP2K4 | 14296 | -1.233 | -0.4587 | No | ||
| 152 | CAMK4 | 14327 | -1.247 | -0.4586 | No | ||
| 153 | SFN | 14447 | -1.307 | -0.4632 | No | ||
| 154 | CSNK2A1 | 14622 | -1.386 | -0.4707 | No | ||
| 155 | DAXX | 14635 | -1.395 | -0.4694 | No | ||
| 156 | CTLA4 | 14652 | -1.403 | -0.4683 | No | ||
| 157 | DDIT3 | 14712 | -1.439 | -0.4694 | No | ||
| 158 | YWHAB | 14792 | -1.481 | -0.4717 | No | ||
| 159 | CSNK1A1 | 14797 | -1.485 | -0.4698 | No | ||
| 160 | DUSP1 | 14879 | -1.529 | -0.4720 | No | ||
| 161 | RALB | 14911 | -1.551 | -0.4715 | No | ||
| 162 | SLC9A1 | 14952 | -1.574 | -0.4715 | No | ||
| 163 | PAK1 | 15056 | -1.632 | -0.4748 | No | ||
| 164 | RALA | 15090 | -1.655 | -0.4742 | No | ||
| 165 | MITF | 15122 | -1.675 | -0.4735 | No | ||
| 166 | RPS6KB1 | 15174 | -1.716 | -0.4739 | No | ||
| 167 | MAX | 15243 | -1.769 | -0.4751 | No | ||
| 168 | MKNK1 | 15297 | -1.800 | -0.4754 | No | ||
| 169 | UBE2I | 15301 | -1.803 | -0.4730 | No | ||
| 170 | ATM | 15302 | -1.803 | -0.4705 | No | ||
| 171 | SMAD3 | 15332 | -1.826 | -0.4694 | No | ||
| 172 | E2F5 | 15561 | -1.994 | -0.4790 | No | ||
| 173 | LCK | 15642 | -2.053 | -0.4805 | Yes | ||
| 174 | YWHAG | 15675 | -2.080 | -0.4793 | Yes | ||
| 175 | CBLB | 15678 | -2.083 | -0.4764 | Yes | ||
| 176 | CREBBP | 15702 | -2.106 | -0.4747 | Yes | ||
| 177 | MAP3K7IP2 | 15735 | -2.133 | -0.4734 | Yes | ||
| 178 | KPNB1 | 15749 | -2.141 | -0.4711 | Yes | ||
| 179 | NCOR1 | 15773 | -2.164 | -0.4693 | Yes | ||
| 180 | PIAS4 | 15885 | -2.263 | -0.4721 | Yes | ||
| 181 | RAF1 | 16017 | -2.394 | -0.4759 | Yes | ||
| 182 | IRF4 | 16037 | -2.419 | -0.4735 | Yes | ||
| 183 | HDAC2 | 16088 | -2.468 | -0.4727 | Yes | ||
| 184 | HDAC1 | 16133 | -2.512 | -0.4715 | Yes | ||
| 185 | PTPRK | 16140 | -2.524 | -0.4683 | Yes | ||
| 186 | CREB1 | 16167 | -2.549 | -0.4661 | Yes | ||
| 187 | CALM2 | 16239 | -2.624 | -0.4663 | Yes | ||
| 188 | RBL1 | 16274 | -2.655 | -0.4643 | Yes | ||
| 189 | GATA3 | 16277 | -2.658 | -0.4607 | Yes | ||
| 190 | PML | 16283 | -2.663 | -0.4572 | Yes | ||
| 191 | KPNA2 | 16310 | -2.686 | -0.4548 | Yes | ||
| 192 | CTDSPL | 16334 | -2.711 | -0.4522 | Yes | ||
| 193 | FKBP1A | 16523 | -2.958 | -0.4583 | Yes | ||
| 194 | SAP30 | 16538 | -2.975 | -0.4548 | Yes | ||
| 195 | HSPA8 | 16541 | -2.979 | -0.4507 | Yes | ||
| 196 | MAPK8 | 16556 | -2.993 | -0.4472 | Yes | ||
| 197 | PAK2 | 16651 | -3.116 | -0.4479 | Yes | ||
| 198 | MAPK14 | 16698 | -3.190 | -0.4459 | Yes | ||
| 199 | JUNB | 16800 | -3.351 | -0.4467 | Yes | ||
| 200 | SIN3B | 16802 | -3.352 | -0.4420 | Yes | ||
| 201 | TGFBR1 | 16864 | -3.455 | -0.4404 | Yes | ||
| 202 | POU2F1 | 16867 | -3.457 | -0.4356 | Yes | ||
| 203 | SMURF2 | 16913 | -3.523 | -0.4331 | Yes | ||
| 204 | NCOA2 | 16939 | -3.555 | -0.4294 | Yes | ||
| 205 | CBFB | 16989 | -3.629 | -0.4269 | Yes | ||
| 206 | MAPK1 | 17133 | -3.886 | -0.4292 | Yes | ||
| 207 | SRF | 17174 | -3.958 | -0.4258 | Yes | ||
| 208 | YWHAH | 17289 | -4.152 | -0.4261 | Yes | ||
| 209 | TRAF2 | 17307 | -4.186 | -0.4211 | Yes | ||
| 210 | SIN3A | 17309 | -4.192 | -0.4152 | Yes | ||
| 211 | MAP3K8 | 17320 | -4.218 | -0.4098 | Yes | ||
| 212 | PPM1A | 17322 | -4.226 | -0.4038 | Yes | ||
| 213 | MAP3K3 | 17391 | -4.397 | -0.4013 | Yes | ||
| 214 | RUNX1 | 17402 | -4.421 | -0.3956 | Yes | ||
| 215 | FYN | 17414 | -4.454 | -0.3899 | Yes | ||
| 216 | ATF2 | 17460 | -4.560 | -0.3859 | Yes | ||
| 217 | PIM1 | 17464 | -4.569 | -0.3796 | Yes | ||
| 218 | CALM3 | 17473 | -4.587 | -0.3735 | Yes | ||
| 219 | GSK3B | 17591 | -4.835 | -0.3730 | Yes | ||
| 220 | FASLG | 17629 | -4.926 | -0.3681 | Yes | ||
| 221 | SP1 | 17636 | -4.943 | -0.3614 | Yes | ||
| 222 | PRKCQ | 17644 | -4.969 | -0.3547 | Yes | ||
| 223 | MEF2A | 17651 | -5.007 | -0.3480 | Yes | ||
| 224 | CDK2 | 17662 | -5.028 | -0.3414 | Yes | ||
| 225 | SMAD2 | 17670 | -5.039 | -0.3346 | Yes | ||
| 226 | ITCH | 17735 | -5.234 | -0.3307 | Yes | ||
| 227 | LSP1 | 17753 | -5.292 | -0.3241 | Yes | ||
| 228 | MAP2K6 | 17779 | -5.369 | -0.3179 | Yes | ||
| 229 | TRAF6 | 17804 | -5.432 | -0.3115 | Yes | ||
| 230 | MAP3K1 | 17806 | -5.436 | -0.3038 | Yes | ||
| 231 | RNF111 | 17847 | -5.551 | -0.2981 | Yes | ||
| 232 | PPP2CA | 17858 | -5.604 | -0.2908 | Yes | ||
| 233 | PPP2CB | 17874 | -5.668 | -0.2835 | Yes | ||
| 234 | CCM2 | 17885 | -5.688 | -0.2760 | Yes | ||
| 235 | MAP3K12 | 17894 | -5.720 | -0.2684 | Yes | ||
| 236 | FOXP3 | 17901 | -5.743 | -0.2605 | Yes | ||
| 237 | MAPK9 | 17946 | -5.900 | -0.2546 | Yes | ||
| 238 | NCOA1 | 17974 | -6.035 | -0.2475 | Yes | ||
| 239 | RHOA | 17985 | -6.067 | -0.2395 | Yes | ||
| 240 | MAPK11 | 18122 | -6.671 | -0.2374 | Yes | ||
| 241 | EGR1 | 18190 | -7.067 | -0.2310 | Yes | ||
| 242 | TGFBR2 | 18294 | -7.848 | -0.2255 | Yes | ||
| 243 | NFATC1 | 18298 | -7.855 | -0.2146 | Yes | ||
| 244 | NFATC3 | 18304 | -7.953 | -0.2036 | Yes | ||
| 245 | PDPK1 | 18310 | -8.013 | -0.1925 | Yes | ||
| 246 | SMAD4 | 18320 | -8.095 | -0.1815 | Yes | ||
| 247 | TGFBR3 | 18368 | -8.522 | -0.1720 | Yes | ||
| 248 | AXIN1 | 18371 | -8.558 | -0.1600 | Yes | ||
| 249 | GDI1 | 18384 | -8.666 | -0.1484 | Yes | ||
| 250 | CTDSP2 | 18401 | -8.819 | -0.1367 | Yes | ||
| 251 | GADD45A | 18439 | -9.374 | -0.1255 | Yes | ||
| 252 | HBP1 | 18488 | -10.231 | -0.1136 | Yes | ||
| 253 | DUSP16 | 18511 | -10.949 | -0.0993 | Yes | ||
| 254 | DGKA | 18512 | -10.953 | -0.0838 | Yes | ||
| 255 | BCL2 | 18517 | -11.183 | -0.0681 | Yes | ||
| 256 | NEDD4L | 18518 | -11.213 | -0.0522 | Yes | ||
| 257 | LAMC1 | 18559 | -13.236 | -0.0357 | Yes | ||
| 258 | EGR2 | 18610 | -27.324 | 0.0003 | Yes |